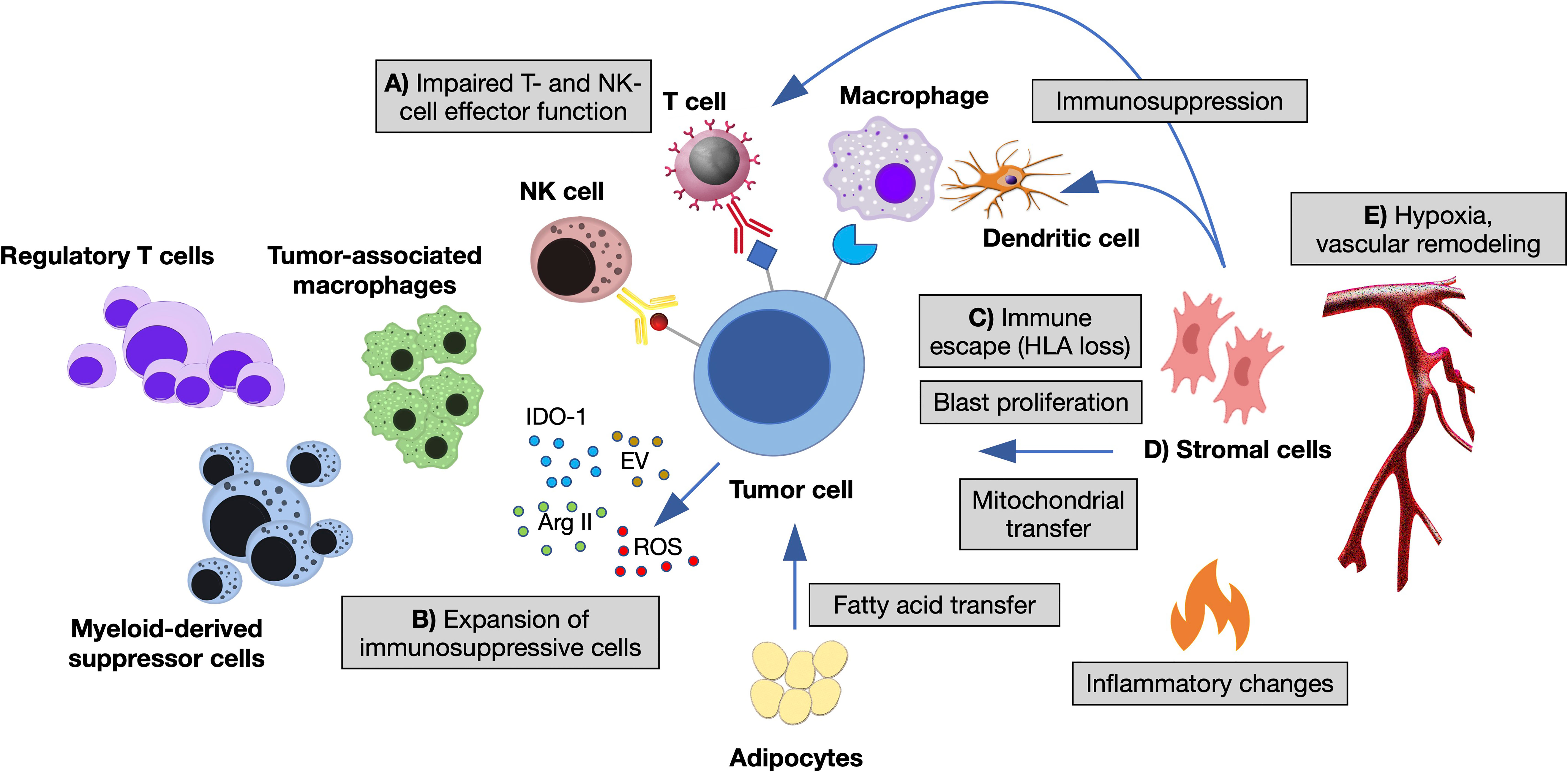
Frontiers | Single-Cell Technologies to Decipher the Immune Microenvironment in Myeloid Neoplasms: Perspectives and Opportunities

Discovery of Novel Allosteric EGFR L858R Inhibitors for the Treatment of Non-Small-Cell Lung Cancer as a Single Agent or in Combination with Osimertinib | Journal of Medicinal Chemistry
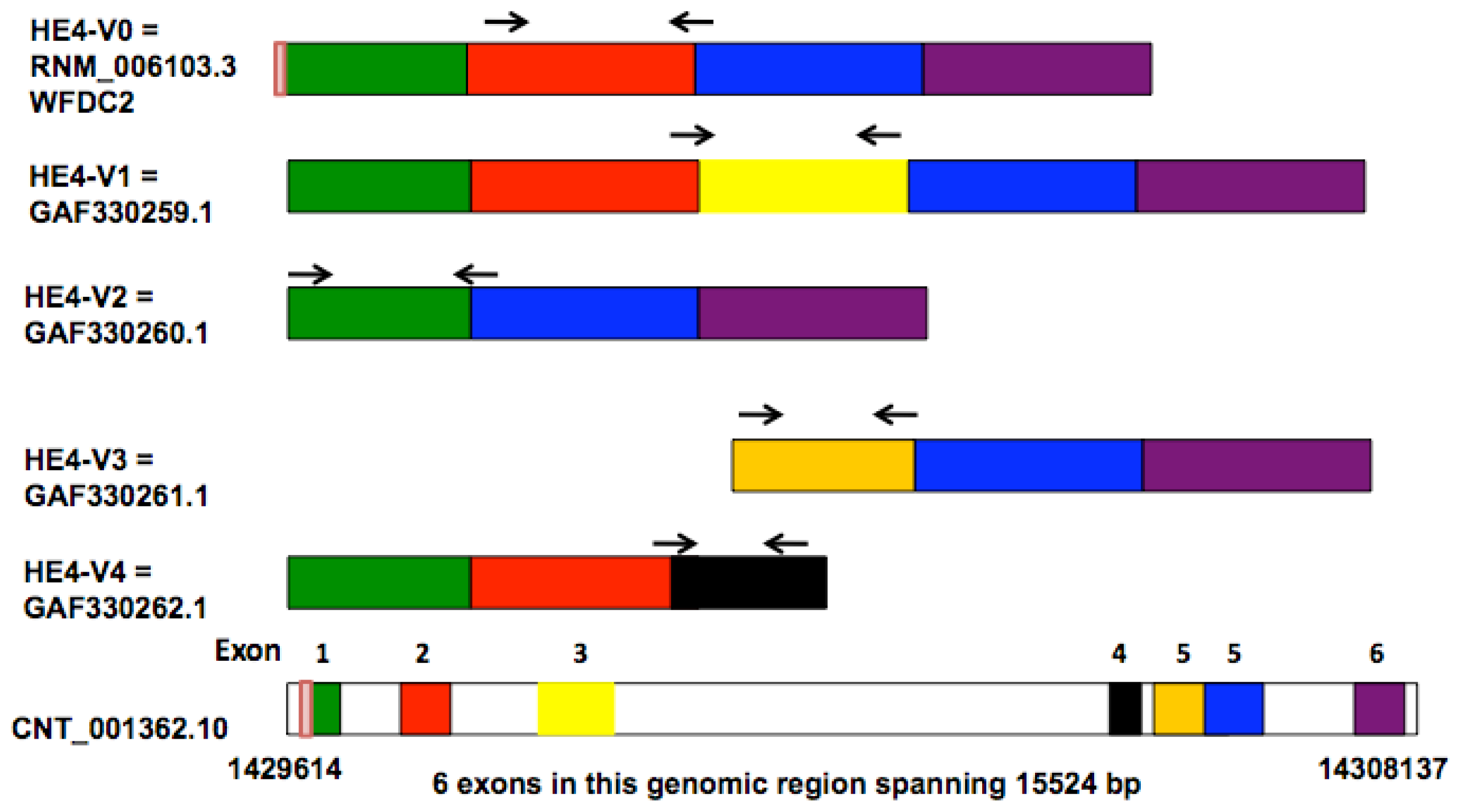
IJMS | Free Full-Text | HE4 Transcription- and Splice Variants-Specific Expression in Endometrial Cancer and Correlation with Patient Survival

Integrative (epi) Genomic Analysis to Predict Response to Androgen-Deprivation Therapy in Prostate Cancer - eBioMedicine
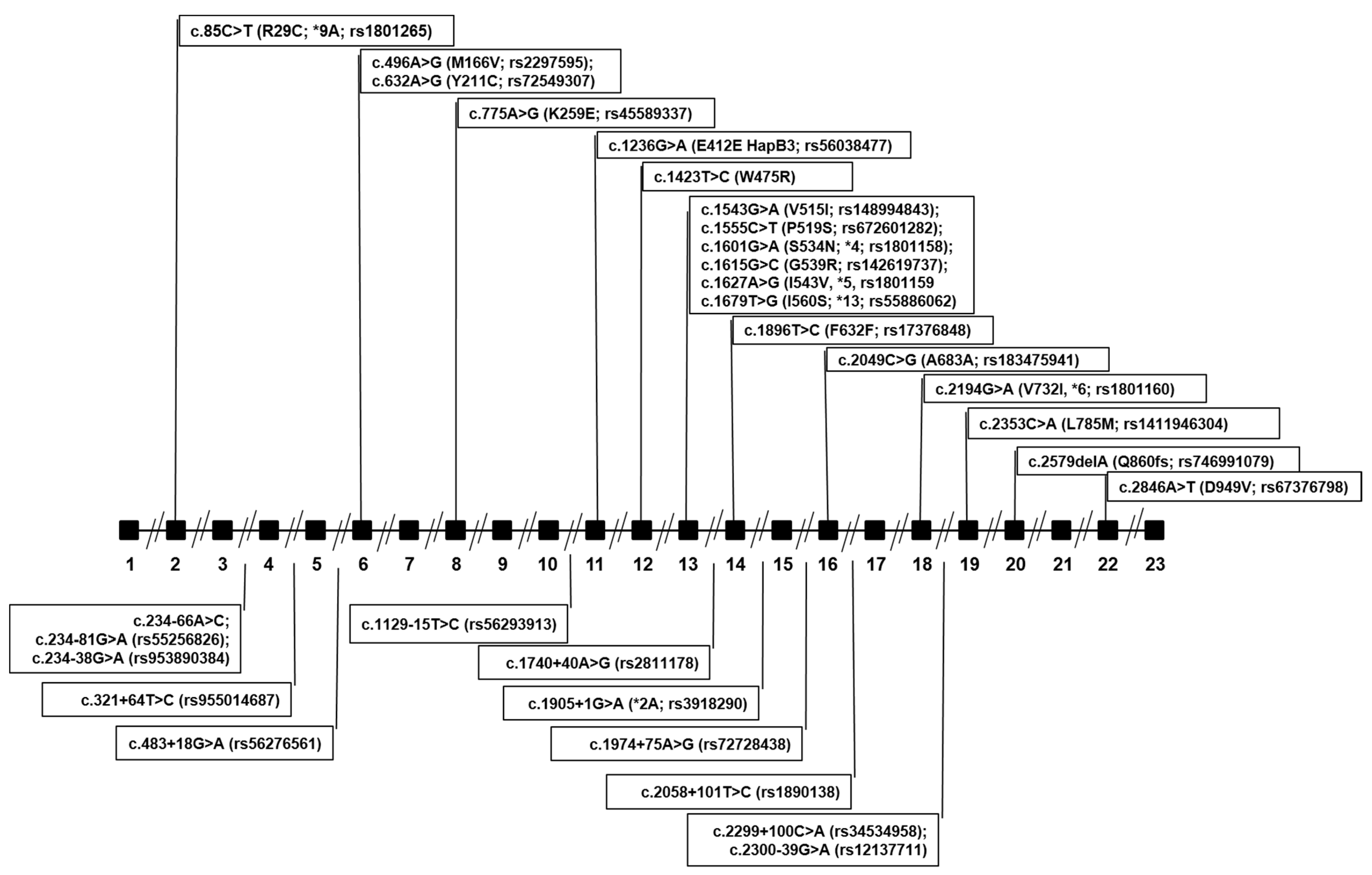
IJMS | Free Full-Text | Predicting Dihydropyrimidine Dehydrogenase Deficiency and Related 5-Fluorouracil Toxicity: Opportunities and Challenges of DPYD Exon Sequencing and the Role of Phenotyping Assays

Dynamic analyses of alternative polyadenylation from RNA-seq reveal a 3′-UTR landscape across seven tumour types | Nature Communications

To nominate potential fusion transcripts, we build a graph from the... | Download Scientific Diagram
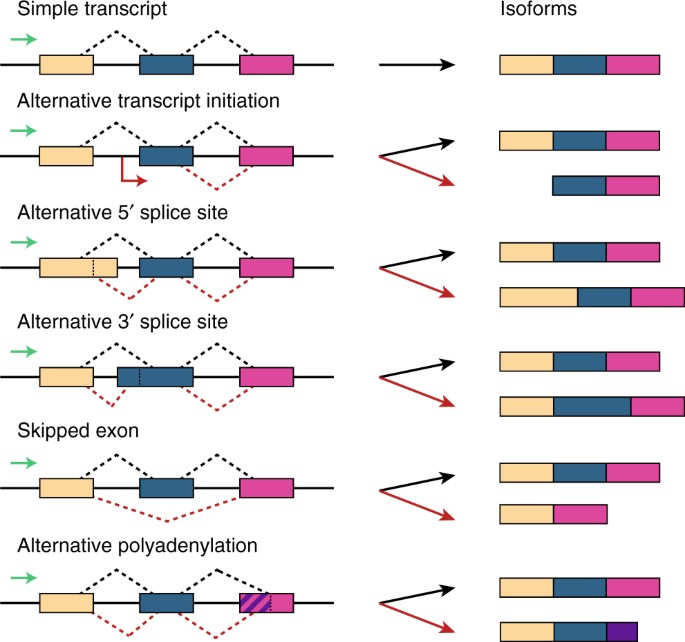
Bayesian nonparametric discovery of isoforms and individual specific quantification | Nature Communications

LongGF: computational algorithm and software tool for fast and accurate detection of gene fusions by long-read transcriptome sequencing | BMC Genomics | Full Text

Optimized Lentiviral Vectors for HIV Gene Therapy: Multiplexed Expression of Small RNAs and Inclusion of MGMTP140K Drug Resistance Gene: Molecular Therapy
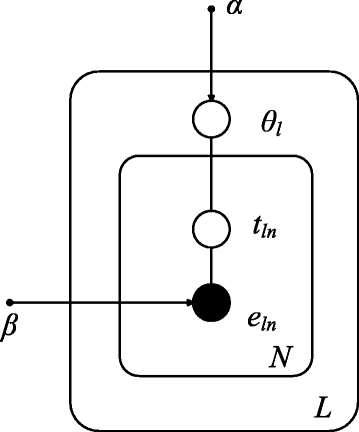
Improving RNA-Seq expression estimation by modeling isoform- and exon-specific read sequencing rate | BMC Bioinformatics | Full Text

The role of brain-derived neurotrophic factor in neural circuit development and function - ScienceDirect

Genes | Free Full-Text | Alternative Splicing Events as Indicators for the Prognosis of Uveal Melanoma
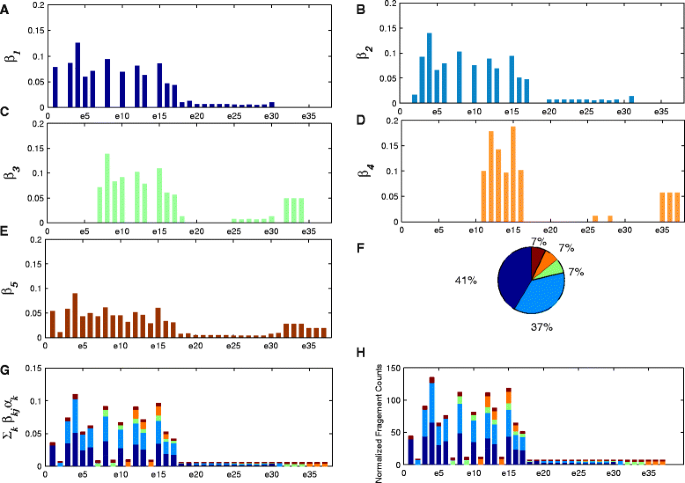
Improving RNA-Seq expression estimation by modeling isoform- and exon-specific read sequencing rate | BMC Bioinformatics | Full Text





